Project Introduction—Second-Prize Solution for MindSpore-based Kidney-Tumor Segmentation
Project Introduction—Second-Prize Solution for MindSpore-based Kidney-Tumor Segmentation
The 10th CCF Big Data & Computing Intelligence Contest (2022 CCF BDCI) has recently come to a close with great success. The winning teams' solutions will be shared on the official competition platform, DataFountain (DF), for further communication and discussion. This blog presents one of the solutions for MindSpore-based Kidney-Tumor Segmentation, which won second prize in the contest. For details about the contest, visit http://go.datafountain.cn/3056.
Team Introduction
Team Name: 大山中学摸奖
Team Members: This team is established by three junior students from the School of Intelligent Systems Engineering, Sun Yan-Sen University. Adhering to the university motto, the team studies extensively on AI theories and technologies and practices earnestly in cutting-edge AI applications. The team also participated in another Huawei's project "SDK-based Person Detection", and won the golden prize related to MindSpore-based project in the 2022 Ascend AI Innovation Contest.
Award: Second Prize
Abstract
The incidence of kidney cancer has been increasing in recent years. How to detect and treat kidney tumors in time becomes a very important issue. Over the last few decades, there has been a significant advancement in deep learning and computer vision technologies, leading to noteworthy breakthroughs in various fields like image classification, object detection, and semantic segmentation. Along with this progress, AI has also been successfully applied in the medical domain, particularly in large-scale and high-accuracy kidney-tumor segmentation.
Despite these achievements, image segmentation presents several challenges that stem from factors such as slice thickness, tumor size, multiple foci, and subtypes of tumors. Given this context, we proposed a hybrid U-Net network based on multimodal data, which uses multimodal inputs to converge 2D information with 3D information. During encoding, an edge-oriented module is introduced to provide fine-grained constraints and edge-related features. During decoding, a large number of residual connections are used to capture fine- and coarse-grained semantics in a full scale, and a weight aggregation module is used to aggregate full-scale information and edge information.
The model we proposed finally got 95.8% (kidney segmentation) and 71.3% (tumor segmentation) Dice scores on the KiTS19 dataset. While compared to other methods, our approach yields relatively high accuracy and holds great potential for practical applications, there are still areas where we can further improve its performance.
Keywords
Multimodal data convergence, edge-oriented module, full-scale information, weight aggregation module
1. Overview
In China, the incidence of kidney tumors has increased at an average annual rate of about 6%, with a cumulative increase of 111.72% over the past 20 years. It can be foreseen that in the near future, kidney tumors will become another "killer" that seriously threatens people's health, especially men's health.
Traditionally, nephrectomy is performed to treat this disease. But to protect kidney functions, excision of only infected kidney parts has recently become a standard treatment for kidney tumors. Clinical practitioners primarily rely on imaging techniques like CT scans to assess treatment options, which encompass total excision, partial excision, and active monitoring. This process involves a substantial amount of manual effort and may also yield inconsistent results between experts who score solutions.
Hence, there has been a growing interest in developing intelligent automated systems for evaluating kidney tumors that can overcome the limitations associated with manual scoring. We expect such a system is able to automatically identify the kidney and tumor regions in medical images to determine whether tumors are benign or malignant and to evaluate the invasiveness of tumors. The automatic segmentation of the kidney and tumor regions is a fundamental function among these, which greatly assists inmedical diagnosis.
Over the past few years, AI has found widespread application in the field of imaging, with a succession of neural network-based methods being developed for tasks such as image classification, image segmentation, and object detection. Traditional CNN networks do not perform well in predicting outcomes for medical images due to their high resolution, multimodal nature, single pixel features, blurred edge shapes, and unbalanced category distribution. Therefore, we improved the U-Net network structure and designed a hybrid U-Net network based on multimodal data. The 2D information and 3D information can be input to our network at the same time, and are converged in the encoding process. The edge information of images is extracted, and the accuracy of image segmentation can be improved according to the constraints of the edge. Besides, a large number of residual connections are used for the decoder to integrate the information at each scale, and weight aggregation is performed to filter the multi-scale information to retain valuable information. In the end, images are segmented based on the combined edge information and multi-scale information after weight aggregation. Experiments have shown that our method has an excellent performance in kidney-tumor segmentation, and there is significant potential for further improvement.
The innovations of our method are as follows:
(1) Use multimodal inputs to combine 2D information with 3D information effectively.
(2) Use the 3D convolution's capability of extracting spatial features, and avoid large memory usage caused by the 3D convolutional network.
(3) Extract and utilize the edge information through multiple modules, and enhance the anti-aliasing effect of image segmentation.
(4) Use a small model. The final weight file size of the model is only 12 MB, and the inference time of a single step is only 81.58 ms, which could even be further improved and has great values in practical projects.
2. Literature Review
In 2015, Olaf Ronneberger and other researchers proposed the U-Net network for biomedical image segmentation and cell segmentation tasks[1]. Since then, U-Net has become one of the most commonly used segmentation models, due to its simple structure, high efficiency, easy to understand and construct, and high accuracy. Generally, a shallow network in a CNN focuses on detailed texture features, and a deep network focuses on semantic features. Both shallow and deep features have their specific meanings. For a segmentation task for medical images, both semantic features and details are required to retrieve the edge features lost in the downsampling process, which is achieved by combining low-layer and high-layer features. This is why U-Net can achieve a good performance.
In 2019, Huazhu Fu proposed the ET-Net model[2], which innovatively introduces the Edge Guidance Module (EGM) and Weighted Aggregation Module (WAM) for edge information extraction and multi-scale information aggregation, respectively.
At the same year, Yifei Zhang proposed the CMnet model[3], which creatively improves the model performance through multimodal data convergence. The multimodal data convergence refers to a process where feature representations of the same example obtained through various means are input into a network, so that the model could have a more comprehensive understanding of the sample features.
In 2020, Lanfen Lin presented the U-Net3+ model[4], which uses a large number of residual connections. Additionally, each decoder integrates small-scale, same-scale, and large-scale feature maps, facilitating information exploration in multiple scales. This significantly improves the accuracy and efficiency of biomedical image segmentation.
3. Analysis of the Contest and Data
3.1 Contest Requirements
The goal of this contest is to develop a fast and reliable semantic segmentation method for kidneys and tumors. The biggest challenge lies in dealing with the morphological heterogeneity and fuzzy tissue boundaries. The contest provides abdominal CT scan data in the arterial phase from 210 patients who have underwent partial or radical nephrectomy and marks the actual semantic segmentation results.
Participants are expected to develop algorithm models that can be used for semantic segmentation of kidneys and tumors, and design kidney-tumor segmentation algorithms based on high-performance deep learning models for fast and accurate segmentation of tumors. The one whose model obtains the most accurate results for semantic segmentation of kidneys and tumors is the winner.
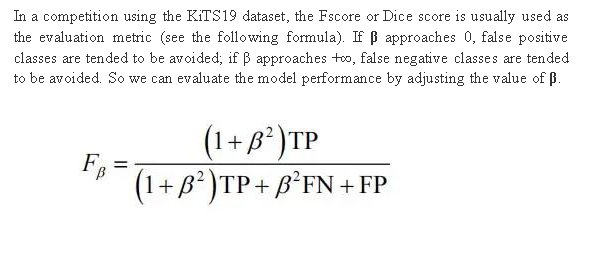
3.2 Evaluation Metric
It's hard to distinguish a false positive class (healthy tissue regarded as tumors) or a false negative class (tumors regarded as healthy tissue), without much reference. In clinical surgery, false negative classes may be considered more serious because they may cause tumors to be partially removed, but tumor tissue is different from a healthy kidney. This is not an urgent concern for doctors. By contrast, some people may think false positive classes are more serious, because they may cause overtreatment on kidney tumors.

3.3 Data Analysis
3.3.1 Complexity of CT Images
The provided CT scan data of 210 cases has different slice thicknesses. As a result, it may be difficult for a model to understand 3D data features when slices are distant. In addition, if the change range of slice thickness is large, it directly leads to anisotropy of data in a depth direction and mainly affects the 3D convolutional layer that cannot understand spatial anisotropy information.
Furthermore, the length of Field of View (FOV) is different. A long FOV length usually leads to a large range of anatomic features, even including images from the top of the head to the bottom of the foot in some cases. As a result, models trained entirely by abdominal scan data may cause false positive classes in unfamiliar areas.
3.3.2 Complexity of Tumor Data
In the data provided, some samples have multiple lesions, which is rare. This could lead network models to tend to predict a single tumor region. In most samples, the size of the tumor accounts for only a small portion of a CT image. A smaller tumor indicates a higher proportion on the regional boundary. However, most segmentation errors occur on the region boundary, resulting in a low Dice score. In addition, smaller tumors are usually more likely to be completely ignored. Kidney tumors are not just manifested as one form, but have many subtypes. Different kidney tumor subtypes have different typical CT imaging manifestations, and some subtypes may be easier to identify than others. While some subtypes, such as angiomyolipoma, have low attenuation, making it difficult to distinguish them from renal cysts. In a word, tumors have individual differences and present diversified features in forms, texture, and grayscale distribution in CT images.
4. Hybrid U-Net Based on Multimodal Data
4.1 Network Structure
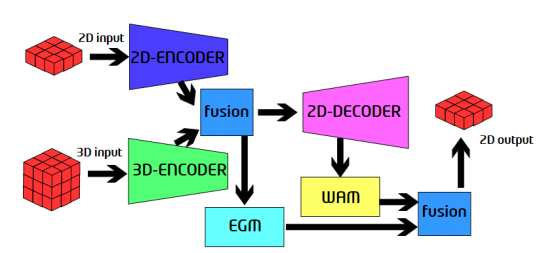
Figure 1. Proposed network structure
This network has two inputs, 2D data and 3D data. The data is passed into the 2D encoder and 3D encoder. After encoding, outputs of the 2D encoder and 3D encoder are combined, and then the EGM module is used to extract the edge information of the features with high resolution. The fusion module connects two tensors on a channel, and then performs convolution to obtain a fusion result. The outputs of the encoders are fused and then passed to the decoder. Then, the WAM module is used to extract the full-scale information, and the information is combined with the outputs of the EGM module to obtain the final prediction result. The structures of the encoder and decoder are designed based on U-Net3+[4], and the detailed description is not presented here.
4.2 3D Encoder
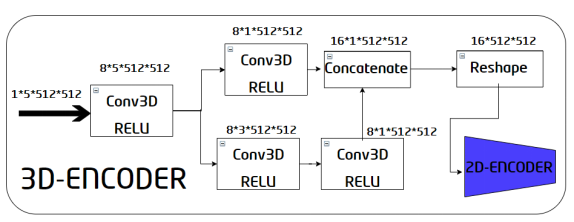
Figure 2. Internal structure of the 3D encoder
The 3D encoder consists of a converter and a 2D encoder. The inputs of the converter are 3D image inputs. First, a 3D convolution is performed to preliminarily obtain a feature map, and then the obtained feature map is processed by using two paths.
The first path is a 3D convolution with a kernel size of 5 x 5 x 5, and the second path contains two 3D convolutions with a size of 3 x 3 x 3. The two paths are used to capture information from areas with different kernel sizes and enrich each other's outputs. Then, combine the outputs of each path to obtain a feature map whose size is 16 x 1 x 512 x 512, and then reshape it into a 16 x 512 x 512 feature map. The feature map of this converter becomes the input of the 2D encoder. During this process, the 3D convolution is used to extract spatial information, and dimensions are reduced through convolutions. This enables the 2D-encoder, thereby avoiding large network memory occupation.
4.3 Edge Guidance Module
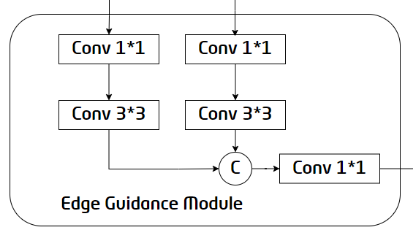
Figure 3. Internal structure of the EGM
The edge information provides useful fine-grained constraints for feature extraction during segmentation, so introducing edge-related features into segmentation tasks can improve segmentation performance. But this requires the feature maps extracted by the first two encoders to have high resolution. Therefore, the EGM selects the feature maps output by the E-BLock1 and E-BLock2 as the inputs. The E1 feature map and the upsampled E2 feature map are combined after 1x1 and 3x3 convolution, and then 1x1 convolution is performed to generate the final edge detection result, providing useful edge features for the subsequent decoder.
4.4 Weighted Aggregation Module
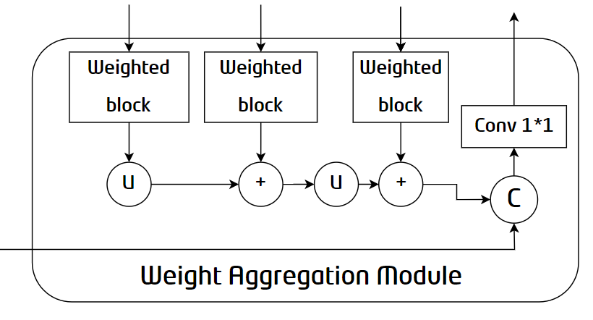
Figure 4. Internal structure of the WAM
To adapt to the shape and size changes of objects, existing methods tend to summarize multi-scale outputs along the channel dimension for final prediction. However, not all channels in the high-level feature map are conducive to object recovery. To solve this problem, WAM, focusing on valuable features, aggregates multi-scale information and edge constraints to improve segmentation performance. As shown in the structure diagram, the network passes the outputs of each D-Block into the Weighted Block to highlight valuable information. Then, the information extracted from each layer is integrated to obtain module outputs.
4.5 Weighted Block
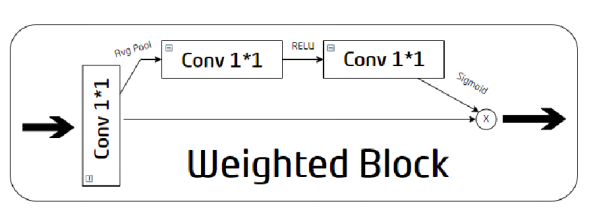

Figure 5. Internal structure of the Weighted Block
The Weighted Block first reconstructs inputs with 1x1 convolution through two paths. The upper path is used to extract weights. First, global average pooling is used to aggregate input global context information, and then two 1×1 convolutional layers with ReLU and Sigmoid are used to estimate correlations. The weights are generated along a channel dimension. No operation is performed on the lower path. The intersection point of the two paths is used to obtain representative features by multiplying the extracted weights with the original modified outputs.
5. Effect
We have continued to fine-tune our model, and during the latest verification, it achieved Dice scores of 0.95819 and 0.71307 for kidney and tumor segmentation respectively. The average Dice score obtained was 0.83563. The following figures show some test effects. (The left part is the CT image, the middle part is the actual segmentation result, and the right part is the model prediction result.)
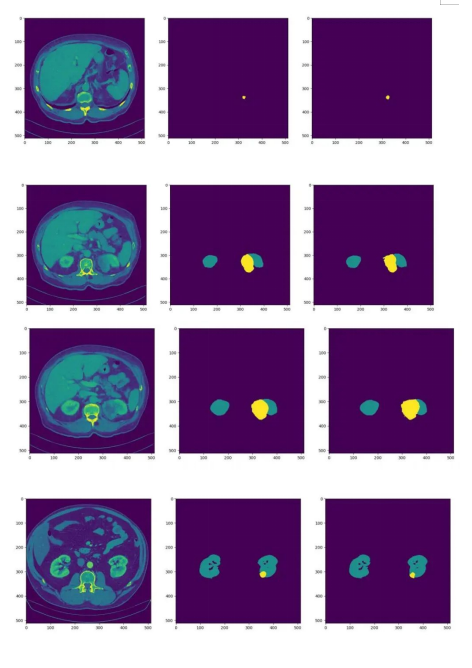
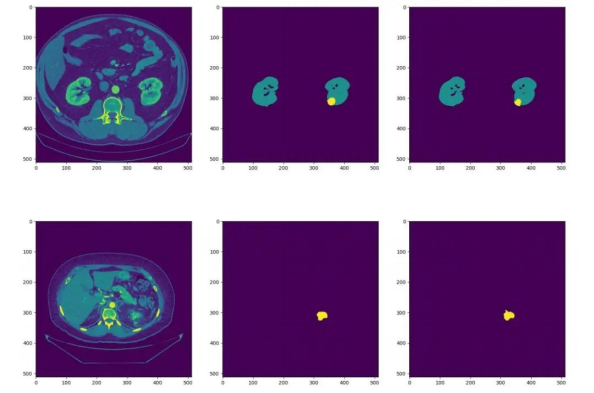
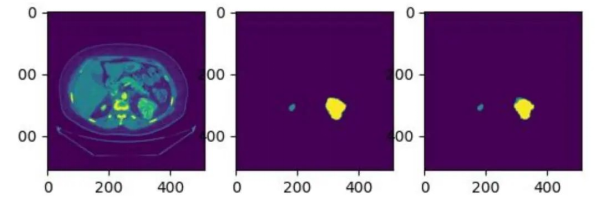
Figure 6. Effect display
6. Conclusion
We innovatively proposed a hybrid U-Net network based on multimodal data. The network inputs are 2D and 3D data, which is encoded through independent encoders. And then the outputs of the 2D and 3D encoders are combined by referring to the combination policy of CMNet[3]. The 3D encoder effectively uses the spatial extraction capability of 3D convolutions, which avoids excessive memory usage. Referring to the ET-Net modules[2] and full-scale connections of U-Net3+[4], we designed modules for edge extraction and weight aggregation, and added a lot of residual connections to the decoder. In addition, we performed upsampling using bilinear interpolation, reducing parameter quantity and network complexity. Our method effectively improves the effect of kidney-tumor segmentation and compresses the network size, making the network easy to train and verify and achieving excellent results on experiments. This provides a feasible solution in the kidney-tumor segmentation field.
Reference
[1]Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation[C]//International Conference on Medical image computing and computer-assisted intervention. Springer, Cham, 2015: 234-241.
[2]Zhang, Z., Fu, H., Dai, H., Shen, J., Pang, Y., Shao, L. (2019). ET-Net: A
Generic Edge-aTtention Guidance Network for Medical Image Segmentation. In: , et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. MICCAI 2019.
[3]Zhang, Yifei et al. “Exploration of Deep Learning-based Multimodal Fusion for Semantic Road Scene Segmentation.” VISIGRAPP (2019).
[4]Huang H, Lin L, Tong R, et al. Unet 3+: A full-scale connected unet for medical image segmentation[C]//ICASSP 2020-2020 IEEE International
Conference on Acoustics, Speech and Signal Processing (ICASSP). IEEE, 2020: 1055-1059.