Project Introduction | MindSpore-based Malaria Detection - Interpretation of Malaria Pathological Sections
Project Introduction | MindSpore-based Malaria Detection - Interpretation of Malaria Pathological Sections
Author: Re Dai Yu Source: Zhihu
Abstract
Malaria is widespread globally, affecting approximately 40% of the world's population. Particularly, it prevails in regions such as Africa, Southeast Asia, Central Asia, and Central and South America. In Africa, malaria is a severe health concern, with around 500 million people residing in malaria-endemic areas. Globally, 100 million individuals are diagnosed with and die of malaria annually, and 90% of these cases occur in Africa.
In many parts of Africa, poverty persists, and inadequate sanitation and medical facilities hinder effective disease identification, including malaria. With the rapid development of artificial intelligence, computer-aided medical image analysis has been booming and supplemented clinical treatments in underdeveloped areas. Leveraging machine learning and deep learning, computer-aided malaria diagnosis is now achievable.
By using thin blood smears from malaria patients as datasets, training models can be developed to determine whether an individual has malaria. Specifically, image analysis is performed on the to-be-tested sample to determine whether this sample belongs to a malaria patient, so as to implement quick diagnosis of malaria. This technology is particularly beneficial in disadvantaged regions as it helps to fill the gaps in local medical resources and personnel, ensuring accurate and efficient malaria diagnosis and assisting in follow-up treatments.
01 Project Design
1.1 Model Principles
1.1.1 Introduction
The model used in this project is Vision Transformer (ViT), a Transformer-based model proposed by the Google's team in 2020 for image classification. ViT has been adopted for a various vision tasks due to its simplicity, efficiency, and scalability, making it a benchmark in the computer vision fields. ViT is ideal for natural language processing and computer vision. It can achieve outstanding performance on image classification tasks without depending on convolution operations.
1.1.2 Model Structure
The main structure of ViT is developed on the encoder part of the Transformer model, but the sequence of some structures is adjusted. For example, the position of Normalization is different from that of the standard Transformer. For details about its structure, see the following figure.
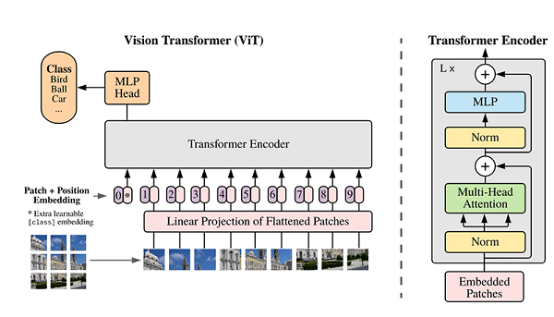
1.1.3 Model Features
ViT is mainly applied to the image classification field. Compared with the conventional Transformer model, it has the following features:
· After a source image of a dataset is divided into a plurality of patches, a two-dimensional patch (without considering channels) is converted into a one-dimensional vector. This vector, a category vector, and a location vector all form a model input.
· Although its block structure is slightly different from that of Transformer, as aforementioned, its main structure is still the Multi-head Attention structure.
· A fully connected layer follows stacked blocks are stacked and accepts the output of the category vector as the input for classification. Generally, the last fully connected layer is called a head, and the Transformer encoder is called backbone.
1.1.4 Model Parsing
The Transformer model is firstly proposed in a 2017-published paper. The encoder-decoder structure based on the Attention mechanism proposed in the paper has achieved great success in the field of natural language processing. The model structure is shown as the following figure.
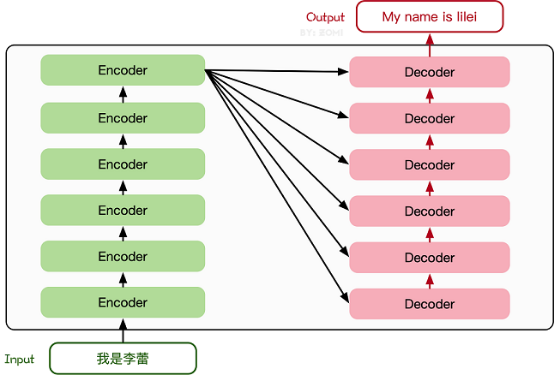
The main structure consists of multiple encoders and decoders. The following figure shows their detailed structures.
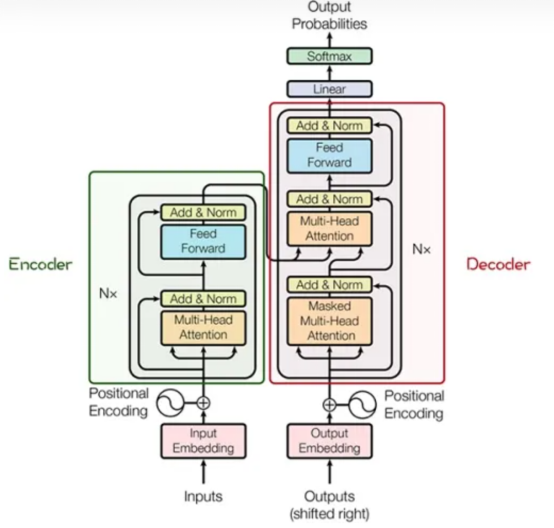
The encoder and decoder consist of many modules, such as the Multi-Head Attention layer, Feed Forward layer, Normalization layer, and Residual Connection ("Add" in the figure). And the most important structure is the Multi-Head Attention structure, which originates from the Self-Attention mechanism and consists of multiple Self-Attentions in parallel.
1.2 Model Training and Effect
1.2.1 Model Training
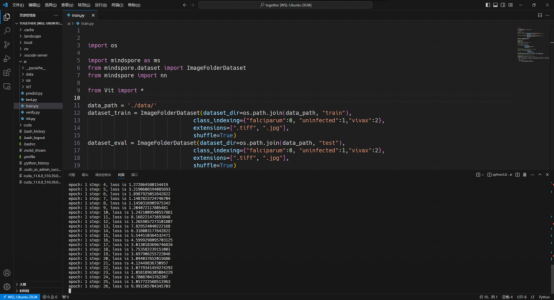
The training dataset used by the model originates from National Library of Medicine. To download the dataset, visit https://lhncbc.nlm.nih.gov/LHC-research/LHC-projects/image-processing/malaria-datasheet.html. After you download the dataset, preprocess it based on the MindSpore dataset. Then, create a ViT model after importing data. For details about the model build process, see the following figure.
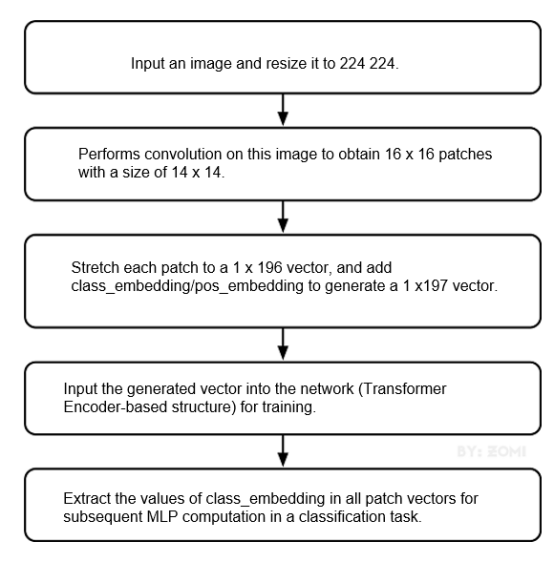
It takes a long time to train a complete ViT model. In actual practices, you are advised to adjust the epoch size based on project requirements. Due to the limited computing power of this project, epoch_size is set to 10, momentum to 0.9, and num_classes to 1000. The following figure shows the training process.
2.2.2 Model Effect Evaluation
1. The Top_1_Accuracy and Top_5_Accuracy evaluation indicators commonly used in the industry are used to evaluate the model performance.
· Top_1_Accuracy measures the accuracy between the category that ranks first and the actual result. That is, the label associated with the highest probability in the final probability vector serves as the prediction result. If the category with the largest probability in the prediction result is correct, the prediction is correct. Otherwise, the prediction is incorrect.
· Top_5_Accuracy measures the accuracy of the top 5 categories that contain the actual result. That is, among the top 5 categories with the largest probability vector, the prediction is correct as long as the correct probability occurs. Otherwise, the prediction is incorrect.
The following lists the model evaluation results of this project.
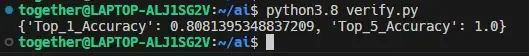
As shown in the preceding figure, Top_1_Accuracy is 0.8081 and Top_5_Accuracy is 1.0, indicating that the model has excellent accuracy.
2. Display of Output Images
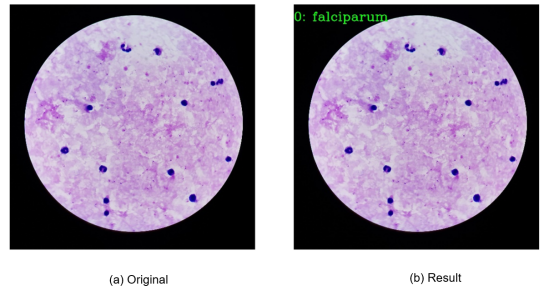
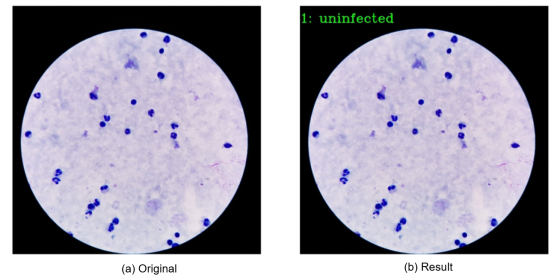
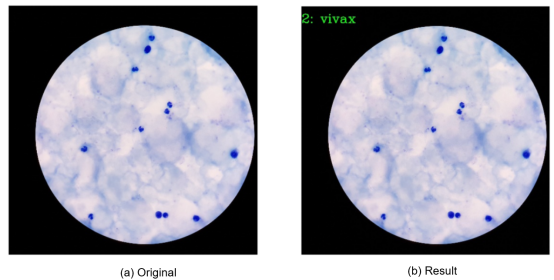
02 MindSpore Installation
Search for a proper computer system version on the MindSpore official website.
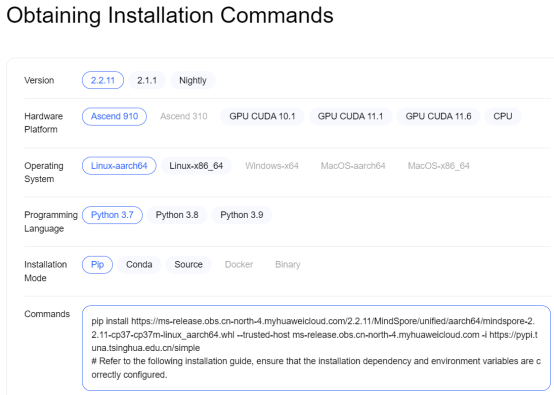
Note that the version used in this project is 2.1.0. The following is the pip installation command:
$pip install https://ms-release.obs.cn-north-4.myhuaweicloud.com/2.1.0/MindSpore/
unified/x86_64/mindspore-2.1.0-cp38-cp38-linux_x86_64.whl --trusted-host
ms-release.obs.cn-north-4.myhuaweicloud.com -i https://pypi.tuna.tsinghua.edu.cn/simple
03 Application of the Project
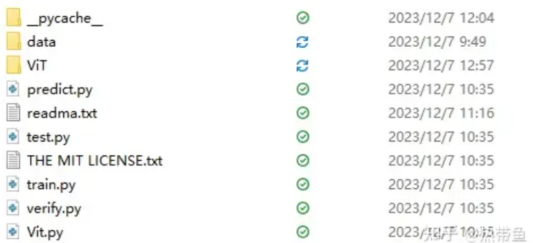
● The structure of this project is shown in the preceding figure.
The datasets, including the test dataset (test) and training dataset (train) are stored in the data folder.
● The ViT folder contains trained models displayed on the web page.
● The python files include a test program, training program, and verification program.
● If you are a beginner, remember to modify the file address before using it. Generally, there is a high probability that an error is reported because the environment configuration is incorrect (i.e. incorrect version) or the file address is incorrectly modified (i.e. address not modified or the slash (/) and backslash (\) not changed).
● Feel free to ask any further questions.
● This project is completely open-source based on the MIT protocol. To be concise, you are allowed to close the source code after it is modified by other developers. You do not need to describe the modified files. And you can use the name of the original author for marketing for further software development.
04 Summary
MindSpore is used to implement data processing, model training, and model inference of this project. Top_1_Accuracy=0.8081 and Top_5_Accuracy=1.0 demonstrate that the model has optimal accuracy. In addition, when an image is input for inference, the prediction result can be viewed.
Through the interpretation of this deep learning-based malaria pathological analysis project, how to build a deep learning-related computer environment and create a ViT model is also elaborated on.
This project can be used on MindSpore-powered hardware developer board for developing inference applications. For details, see the official MindSpore documentation.
References
[1]https://www.mindspore.cn/tutorials/application/en/r2.1/cv/vit.html
[2]https://www.mindspore.cn/tutorials/application/en/r2.0/cv/vit.html