vmap: A Powerful Utility for AI Research
vmap: A Powerful Utility for AI Research
Author: Yeats_Liao | Source: MindSpore Forum
The vigorous development of AI converged computing poses new requirements and challenges to framework capabilities. Problem scenarios and model design become increasingly complex, making the service data dimensions and the nesting depth of computing logic increase accordingly. Even if the vectorization optimization method can effectively resolve performance bottlenecks, it is not easy for common users to implement. It might be easy to implement low-dimensional data operations. However, as the data dimensions increase, the service logic becomes more complex, which requires us to clearly understand the logic mapping between data dimensions of operations, bringing great challenges to model design and coding. The automatic vectorization feature, vmap, helps solve this problem, which allows us to separate specific batch processing logic from functions. When writing a function, we only need to consider the low-dimensional operation logic. The vmap API is called to automatically implement high-dimensional operation. In addition, nested calling is supported, which effectively reduces problem complexity.
This blog describes how to use the vmap API to convert highly repeated operation logic in models or functions into parallel vector operation logic, achieving simplified code logic and efficient execution performance.
01
Environment Setup
Go to the MindSpore official website and click Install in the upper menu bar.
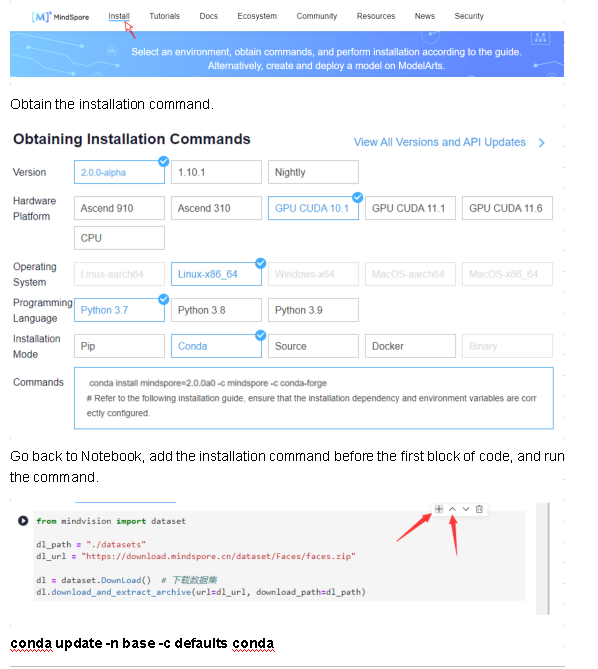
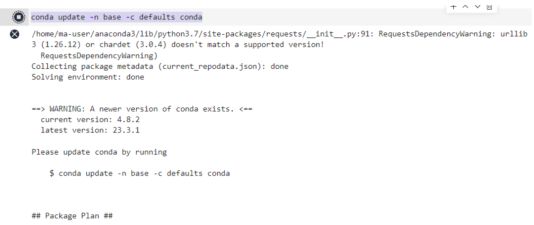
Install the MindSpore 2.0 GPU version.
conda install mindspore=2.0.0a0 -c mindspore -c conda-forge
Install Mindvision.
pip install mindvision
02
Vectorization Thinking
Vectorization thinking is a common technology that improves computing performance. Vectorization thinking can be expressed as follows:
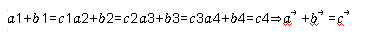 ⃗
⃗
The core idea is to convert the operation logic of multiple for loops into one computation of vectors. Vectorization thinking still works when it comes to a function or a set of operations of a model.
03
Manual Vectorization
First, we construct a simple convolution function, which is applicable to one-dimensional vector scenarios.
Import mindspore
From mindspore import Tensor, ops
Import mindspore.numpy as mnp
x = mnp.arange(5).astype('float32')
w = mnp.array([1., 2., 3.])
def convolve(x, w):
output = []
for i in range(1, len(x) - 1):
output.append(mnp.dot(x[i - 1 : i + 2], w))
return mnp.stack(output)
convolve(x, w)
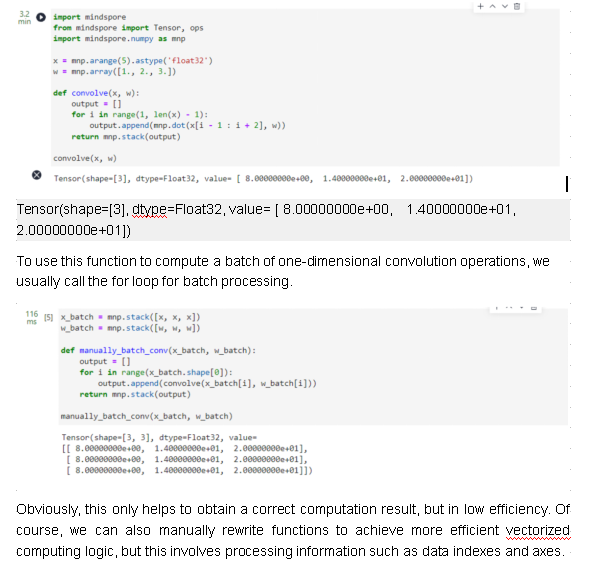
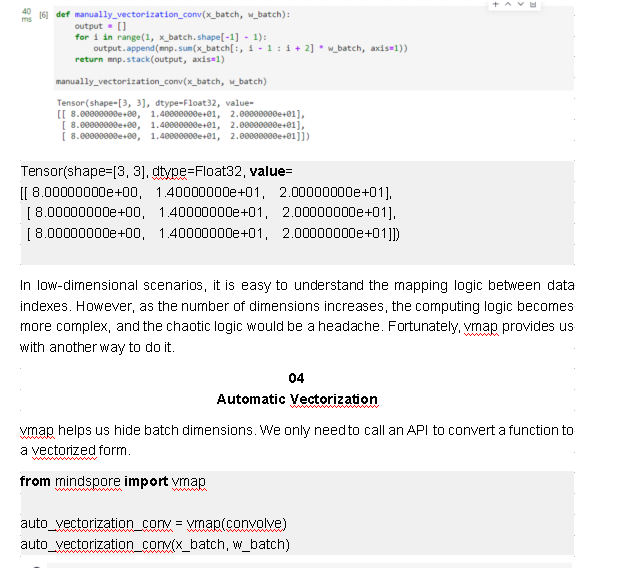
def manually_vectorization_conv(x_batch, w_batch):
output = []
for i in range(1, x_batch.shape[-1] - 1):
output.append(mnp.sum(x_batch[:, i - 1 : i + 2] * w_batch, axis=1))
return mnp.stack(output, axis=1)
manually_vectorization_conv(x_batch, w_batch)
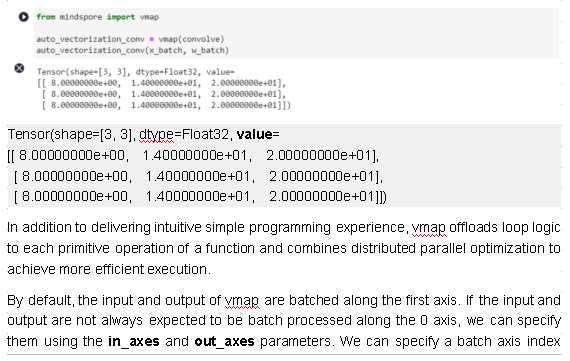
separately for each input or output, or specify the same batch axis index for all inputs or outputs.
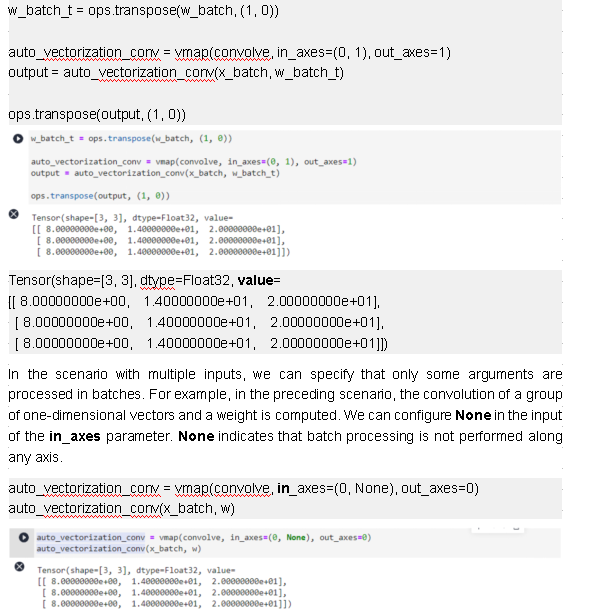
Tensor(shape=[3, 3], dtype=Float32, value=
[[ 8.00000000e+00, 1.40000000e+01, 2.00000000e+01],
[ 8.00000000e+00, 1.40000000e+01, 2.00000000e+01],
[ 8.00000000e+00, 1.40000000e+01, 2.00000000e+01]])
To ensure the correctness of the vmap operation logic, vmap verifies the input dimension, axis index, and batch size. For details about the parameter restrictions, see mindspore.vmap.
Nesting of High-Order Functions
vmap is essentially a high-order function that takes the function as the input and returns a vectorized function that can be applied to batch data processing. It can be nested and combined with high-order functions provided by other frameworks.
The vmap API is nested and applies to the batch processing logic of more than two layers.
hyper_x = Tensor([[1., 2., 3., 4., 5.], [2., 3., 4., 5., 6.], [3., 4., 5., 6., 7.]], mindspore.float32)
hyper_w = Tensor([[1., 1., 1.], [2., 2., 2.], [3., 3., 3.]], mindspore.float32)
hyper_vmap_ger = vmap(vmap(convolve, in_axes=[None, 0]), in_axes=[0, None])
hyper_vmap_ger(hyper_x, hyper_w)
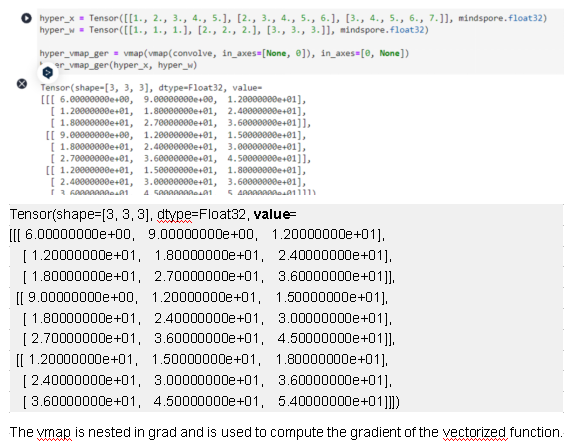
from mindspore import grad
def forward_fn(x, y):
out = x + 2 * y
out = ops.sin(out)
return ops.reduce_sum(out)
x_hat = Tensor([[1., 2., 3.], [2., 3., 4.]], mindspore.float32)
y_hat = Tensor([[2., 3., 4.], [3., 4., 5.]], mindspore.float32)
grad_vmap_ger = grad(vmap(forward_fn), grad_position=(0, 1))
grad_vmap_ger(x_hat, y_hat)
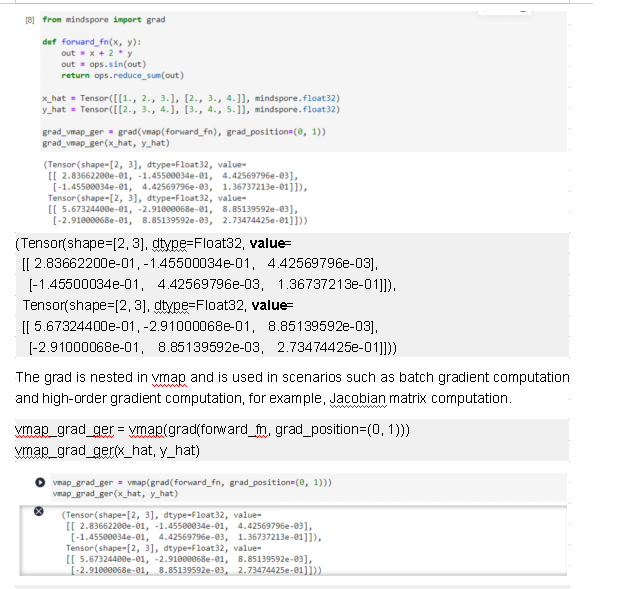
(Tensor(shape=[2, 3], dtype=Float32, value=
[[ 2.83662200e-01, -1.45500034e-01, 4.42569796e-03],
[-1.45500034e-01, 4.42569796e-03, 1.36737213e-01]]),
Tensor(shape=[2, 3], dtype=Float32, value=
[[ 5.67324400e-01, -2.91000068e-01, 8.85139592e-03],
[-2.91000068e-01, 8.85139592e-03, 2.73474425e-01]]))
Automatic Vectorization of the Cell Object
In the previous test cases, the function object is used as the input. The following describes how to use the Cell object as the input of vmap. This is an example of a simply defined fully-connected layer.
import mindspore.nn as nn
from mindspore import Parameter
from mindspore.common.initializer import initializer
class Dense(nn.Cell):
def __init__(self, in_channels, out_channels, weight_init='normal', bias_init='zeros'):
super(Dense, self).__init__()
self.scalar = 1
self.weight = Parameter(initializer(weight_init, [out_channels, in_channels]), name="weight")
self.bias = Parameter(initializer(bias_init, [out_channels]), name="bias")
self.matmul = ops.MatMul(transpose_b=True)
self.bias_add = ops.BiasAdd()
def construct(self, x):
x = self.matmul(x, self.weight)
output = self.bias_add(x, self.bias)
return output
input_a = Tensor([[1, 2, 3], [4, 5, 6]], mindspore.float32)
input_b = Tensor([[2, 3, 4], [5, 6, 7]], mindspore.float32)
input_c = Tensor([[3, 4, 5], [6, 7, 8]], mindspore.float32)
dense_net = Dense(3, 4)
print(dense_net(input_a))
print(dense_net(input_b))
print(dense_net(input_c))
inputs = mnp.stack([input_a, input_b, input_c])
vmap_dense_net = vmap(dense_net)
print(vmap_dense_net(inputs))
[[ 0.0219292 -0.01062493 -0.03378957 -0.02589925]
[ 0.03091274 -0.04968021 -0.08098207 -0.07896652]]
[[ 0.02492371 -0.02364336 -0.0495204 -0.04358834]
[ 0.03390725 -0.06269865 -0.09671289 -0.09665561]]
[[ 0.02791822 -0.03666179 -0.06525123 -0.06127743]
[ 0.03690176 -0.07571708 -0.11244373 -0.1143447 ]]
[[[ 0.0219292 -0.01062493 -0.03378957 -0.02589925]
[ 0.03091274 -0.04968021 -0.08098207 -0.07896652]]
[[ 0.02492371 -0.02364336 -0.0495204 -0.04358834]
[ 0.03390725 -0.06269865 -0.09671289 -0.09665561]]
[[ 0.02791822 -0.03666179 -0.06525123 -0.06127743]
[ 0.03690176 -0.07571708 -0.11244373 -0.1143447 ]]]
The usage of Cell is basically the same as that of function-based automatic vectorization. You only need to replace the first input parameter of vmap with the Cell instance. vmap vectorizes construct for batch data processing. In addition, two Parameter arguments are defined for the initialization function in this test case. The vmap processing of free variables of this type of execution functions is equivalent to using the free variables as arguments and setting in_axes to None.
In this way, batch input can be used for training or inference on the same model. Compared with the existing network model input that supports batch axis input, the batch processing dimension implemented by using vmap is more flexible and is not limited to input formats such as NCHW.
Model Ensembling Scenario
In the model ensembling scenario, prediction results from multiple models are combined. Traditionally, each model is run on certain inputs, and then the prediction results are combined. If you are running models with the same architecture, you can vectorize them with vmap for acceleration.
In this scenario, vectorization of weight data is involved. If the running model is implemented through functional programming, that is, weight parameters are defined outside the model and transferred to the model through arguments, you can directly configure in_axes to perform batch processing. MindSpore provides convenient model definitions in that the weight parameters of most neural network (NN) APIs are internally defined and initialized. This means that the weight parameters in the model cannot be processed in batches in the original vmap API. Therefore, extra workload is required for reconstructing the model to a function that is transferred through arguments. Fortunately, the vmap API of MindSpore has optimized this scenario for you. You only need to transfer multiple running model instances to the vmap API in CellList format. Then the framework can automatically implement batch processing of weight parameters.
The following demonstrate how to use a simple set of CNN models to implement model ensembling inference and training.
class LeNet5(nn.Cell):
"""
LeNet-5 network structure
"""
def __init__(self, num_class=10, num_channel=1):
super(LeNet5, self).__init__()
self.conv1 = nn.Conv2d(num_channel, 6, 5, pad_mode='valid')
self.conv2 = nn.Conv2d(6, 16, 5, pad_mode='valid')
self.fc1 = nn.Dense(16 * 5 * 5, 120)
self.fc2 = nn.Dense(120, 84)
self.fc3 = nn.Dense(84, num_class)
self.relu = nn.ReLU()
self.max_pool2d = nn.MaxPool2d(kernel_size=2, stride=2)
self.flatten = nn.Flatten()
def construct(self, x):
x = self.conv1(x)
x = self.relu(x)
x = self.max_pool2d(x)
x = self.conv2(x)
x = self.relu(x)
x = self.max_pool2d(x)
x = self.flatten(x)
x = self.fc1(x)
x = self.relu(x)
x = self.fc2(x)
x = self.relu(x)
x = self.fc3(x)
return x


[[ 1.48978233e-06, 1.02267529e-06, 1.33801677e-06 ... -1.32894393e-05, 1.36311328e-05, -3.29658405e-06],
[ 1.09956818e-06, -5.06103561e-07, 3.04885953e-06 ... -1.76028752e-05, 1.66466998e-05, -1.17290392e-06],
[ 2.96090502e-06, 1.87074147e-06, 5.76813818e-06 ... -1.09994007e-05, 1.35614964e-05, -2.19983576e-06]],
[[ 6.74323928e-06, -1.03955799e-05, -6.92168396e-06 ... 4.88165415e-06, -5.40378596e-06, 3.09346888e-06],
[ 7.28906161e-06, -1.34921102e-05, -1.00995640e-05 ... 9.44596650e-07, -6.40979761e-06, 1.26146606e-05],
[ 9.43304440e-06, -1.61852931e-05, -1.16265892e-05 ... 5.31926253e-06, -1.28484417e-05, 8.03831313e-07]],
[[-5.51165886e-06, -1.09487860e-06, -6.07781249e-06 ... 7.51453626e-06, -3.29403338e-06, 3.45475746e-06],
[-6.27516283e-06, 1.40756754e-06, -9.18502155e-06 ... 4.16079911e-06, -5.30383022e-06, 5.12517454e-06],
[-6.19608954e-06, 5.12868655e-06, -1.00337056e-05 ... 2.93281119e-07, -6.52256404e-06, 3.62988635e-06]]])
In addition to model ensembling inference, the vmap feature can also be used to implement model ensembling training.
from mindspore.common.parameter import ParameterTuple
class TrainOneStepNet(nn.Cell):
def __init__(self, net, lr):
super(TrainOneStepNet, self).__init__()
self.loss_fn = nn.WithLossCell(net, nn.SoftmaxCrossEntropyWithLogits(sparse=True, reduction='mean'))
self.weight = ParameterTuple(net.trainable_params())
self.adam_optim = nn.Adam(self.weight, learning_rate=lr, use_amsgrad=True)
def construct(self, batch, targets):
loss = self.loss_fn(batch, targets)
grad_weights = grad(self.loss_fn, grad_position=None, weights=self.weight)(batch, targets)
self.adam_optim(grad_weights)
return loss
train_net1 = TrainOneStepNet(net1, lr=1e-2)
train_net2 = TrainOneStepNet(net2, lr=1e-3)
train_net3 = TrainOneStepNet(net3, lr=2e-3)
train_net4 = TrainOneStepNet(net4, lr=3e-3)
train_nets = nn.CellList([train_net1, train_net2, train_net3, train_net4])
model_ensembling_train_one_step = vmap(train_nets)
images = Tensor(mnp.randn(4, 3, 1, 32, 32), mindspore.float32)
labels = Tensor(mnp.randint(1, 10, (4, 3)), mindspore.int32)
for i in range(1, 11):
loss = model_ensembling_train_one_step(images, labels)
print("Step {} - loss: {}".format(i, loss))
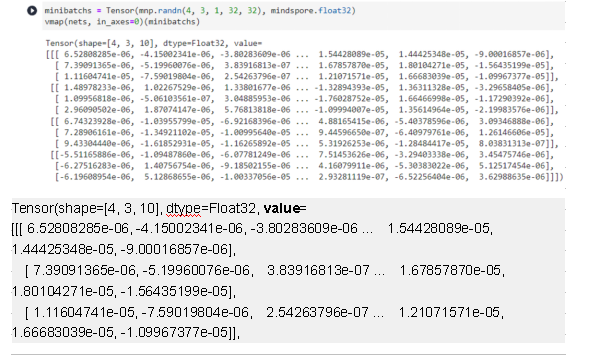
vmap(nets, in_axes=None)(minibatch)
Step 1 - loss: [2.3025837 2.3025882 2.3025858 2.3025842]
Step 2 - loss: [2.260927 2.301028 2.2992857 2.2976868]
Step 3 - loss: [1.8539654 2.2993202 2.2951114 2.2899477]
Step 4 - loss: [0.77165794 2.2973287 2.288719 2.2726345 ]
Step 5 - loss: [0.9397469 2.2948549 2.2777178 2.2313874]
Step 6 - loss: [0.6747699 2.29158 2.2579656 2.1410708]
Step 7 - loss: [0.64673084 2.2870557 2.2232006 1.966973 ]
Step 8 - loss: [1.0506033 2.2806385 2.1645374 1.6848679]
Step 9 - loss: [0.612196 2.2714498 2.0706694 1.3499321]
Step 10 - loss: [0.8843982 2.258316 1.9299208 1.1472267]
Tensor(shape=[4, 3, 10], dtype=Float32, value=
[[[-1.91058636e+01, -1.92182674e+01, 1.06328402e+01 ... -1.87287464e+01, -1.87855473e+01, -2.02504387e+01],
[-1.94767399e+01, -1.95909595e+01, 1.08379564e+01 ... -1.90921249e+01, -1.91503220e+01, -2.06434765e+01],
[-1.96521702e+01, -1.97674465e+01, 1.09355783e+01 ... -1.92643051e+01, -1.93227654e+01, -2.08293762e+01]],
[[-4.07293849e-02, -4.27918807e-02, 5.22112176e-02 ... -4.67570126e-02, -3.88025381e-02, 4.88412194e-02],
[-3.91553082e-02, -4.11494374e-02, 5.00433967e-02 ... -4.48847115e-02, -3.73134986e-02, 4.68519926e-02],
[-3.80369201e-02, -3.99325565e-02, 4.84890938e-02 ... -4.35365662e-02, -3.62745039e-02, 4.54225838e-02]],
[[-5.08784056e-01, -5.05123973e-01, 5.20882547e-01 ... 4.72596169e-01, -5.00697553e-01, -4.60489392e-01],
[-4.80103493e-01, -4.76664037e-01, 4.91507798e-01 ... 4.46062207e-01, -4.72493649e-01, -4.34652239e-01],
[-4.81168061e-01, -4.77702975e-01, 4.92583781e-01 ... 4.47029382e-01, -4.73524809e-01, -4.35579300e-01]],
[[-3.66236401e+00, -3.25362825e+00, 3.51312804e+00 ... 3.77490187e+00, -3.36864424e+00, -3.34358120e+00],
[-3.49160767e+00, -3.10209608e+00, 3.34935308e+00 ... 3.59894991e+00, -3.21167707e+00, -3.18782210e+00],
[-3.57623625e+00, -3.17717075e+00, 3.43059325e+00 ... 3.68615556e+00, -3.28948307e+00, -3.26504302e+00]]])

05
Summary
This blog focuses on the usage of vmap. In essence, vmap does not execute the loop logic outside the function. Instead, it offloads the loop to each primitive operation of the function and transfers the mapping axis information between primitive operations to ensure correct operation logic. The vmap performance benefits mainly come from the VmapRule implementation corresponding to each primitive operation. Because the loop is offloaded to the operator level, it is easier to optimize the performance based on the parallel technology. If you have custom operators in your function, you can try to implement specific VmapRule for custom operators to achieve better performance. If ultimate performance is required, the graph kernel fusion feature can be used for optimization.
Currently, the vmap feature supports the GPU and CPU platforms. More functions are being adapted to the Ascend platform.
If the vmap contains control flows, each batch processing branch must have the same processing operation or all variables in the control flow logic has no split axis.