Project Introduction — Second-Prize Solution for MindSpore-based Kidney-Tumor Segmentation
Project Introduction — Second-Prize Solution for MindSpore-based Kidney-Tumor Segmentation
The 10th CCF Big Data & Computing Intelligence Contest (2022 CCF BDCI) has recently come to a close with great success. The winning teams' solutions will be shared on the official competition platform, DataFountain (DF), for further communication and discussion. This blog presents one of the solutions for MindSpore-based Kidney- Tumor Segmentation, which won second prize in the contest. For details about the contest, visit http://go.datafountain.cn/3056.
Team Introduction
Team Name: E Di Shen Na
Team Members: This team is established by five members from Hunan University, Zhengzhou University, University of Electronic Science and Technology of China, Northwestern Polytechnical University, and Migu Music. With main research focusing on medical image segmentation, image detection, and industry quality inspection, some team members have published related papers and awarded prizes in other Huawei-hosted competitions.
Award: Second Prize
Abstract
Our method is inspired by nnU-Net, a 3D U-Net-based framework that automatically adapts to a given dataset. This framework has the ability to achieve state-of-the-art performance in various tasks without requiring manual parameter adjustments. For our model, we first employ methods, such as cropping, resampling, and normalization to preprocess each training data record. Then, we utilize data argumentation policies, such as random rotation, random scaling, gamma correction, and Gaussian noise. Finally, we execute segmentation with various auxiliaries on the 3D U-Net architecture to supervise the backbone network and surmount issues such as vanishing gradient and sluggish convergence during deep neural network training. In the preliminary round, our model gets a score of 0.8377, ranking second. For details about the code, visit https://github.com/wangwangwang978/MindSpore2022-BDCI.
Keywords
MindSpore, kidney-tumor segmentation, nnU-Net
1 Introduction
Over the past few years, medical image segmentation has seen significant advancements thanks to convolutional neural networks (CNNs). Among these, full convolutional networks (FCNs) have emerged as an ideal solution for performing semantic segmentation on medical images, which greatly assists pathologists in diagnosing patients. U-Net[1], one of its typical variants, consists of a symmetric U-shaped encoder-decoder architecture and several skip connections. The encoder utilizes continuous convolutional layers and downsampling layers to extract deep features. And the decoder upsamples the extracted deep features to the input resolution for pixel-level semantic segmentation, and uses skip connections to fuse high-resolution features of varying scales, mitigating the loss of spatial information that occurs during the downsampling phase of the encoder. Based on this elegant structure, U-Net++ and U-Net3+ networks are proposed. Moreover, 3D U-Net, UNETR, and Swin UNETR have been applied to the medical imaging field, achieving noteworthy segmentation performance[2].
The KiTS19 Challenge dataset is used in the MindSpore-based Kidney-Tumor Segmentation contest, which aims to develop a solution that is reliable for kidney-tumor semantic segmentation. The dataset includes abdominal CT scan data in the arterial phase from 310 patients who have underwent partial or radical nephrectomy, along with corresponding semantic segmentation labels. The data of 210 patients is used for model training and verification, and the remaining data of 90 patients is used for objective model evaluation.
However, according to the previous results of referring to the KiTS19 Grand Challenge and researches of Isensee F[4], simply making changes to the proposed algorithms is not enough to surpass the performance of the optimized 3D U-Net baseline. Therefore, our team adopts nnU-Net[5,6] for this competition.
2 Method
By referring to the open source code of MindSpore and PyTorch, we reproduced the nnU-Net method for MindSpore-based kidney-tumor segmentation on the KiTS19 dataset.
2.1 Preprocessing
2.1.1 Cropping
All training data is cropped to areas with non-zero values. This has no effect on most datasets (such as liver CT data), but reduces the size and computational loads of skull stripping MRI.
2.1.2 Resampling
Different scanners or collection protocols usually result in datasets with heterogeneous voxel spacings. To make the network correctly learn spatial semantics, all CT images are resampled to a unified voxel spacing, where image data uses B-spline interpolation, and the corresponding segmentation mask uses nearest neighbor interpolation. By referring to the champion's solution of KiTS19 challenge[4], we resampled the voxel spacings of all data to 3.22 x 1.62 x 1.62.
2.1.3 Normalization
All intensity values that appear in the training dataset are collected, and the entire dataset is normalized by cropping the intensity values to percentiles in the range of [0.5,99.5]. Then Z-Score normalization is performed based on the average value and standard deviation of all collected intensity values.
2.2 Network Architecture
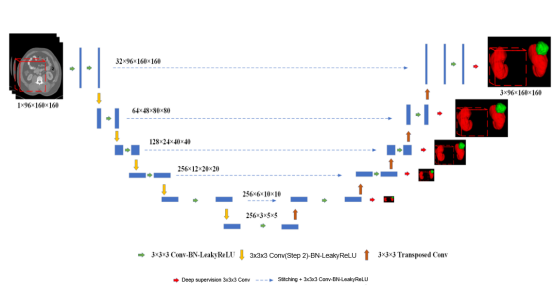
Figure 1 3D U-Net network architecture
In this experiment, we constructed the model architecture based on the U-shaped network structure of 3D U-Net, as shown in Figure 1. The goal was to extract and integrate high-level semantic information and low-level fine features, so as to obtain both context and spatial information. The 3D U-Net network structure consists of an encoder and a decoder. The encoder consists of a 3 x 3 x 3 convolutional layer and a 2 x 2 x 2 maximum pooling downsampling layer. After each downsampling, the number of channels is doubled and the image resolution is halved. The decoder consists of a 2 x 2 x 2 deconvolutional layer, a stitching layer, and a 3 x 3 x 3 convolutional layer, and is finally output after undergoing a 1 x 1 x 1 convolution. After each upsampling, the number of channels is reduced by half and the image resolution is doubled. Through skip connections, features from each layer in the encoder are stitched, and the global context information at high layers and precise details at low layers are integrated. In addition, the loss of spatial information during the downsampling is reduced.
At the same time, the deep supervision output is performed at the resolution of 1, 1/2, 1/4, 1/8, and 1/16 on the decoder. The 1 x 1 x 1 convolutional layer is used to change the number of channels to the number of segmentation categories. This assists in supervising the backbone network to solve problems such as vanishing gradient and slow convergence.
2.3 Loss Function
Our model is trained in an end-to-end manner, and the loss function is used to calculate the weighted Dice loss and cross entropy loss, which is presented as follows:
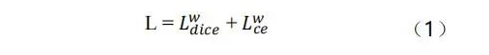
We use deep supervision to improve gradient flows. As shown in the formula (1), segmentation with auxiliaries is added for supervising the backbone network, so as to address the problems of vanishing gradient and slow convergence. The final loss function is presented as the formula (2):

In the formula, w__i indicates a changeable hyperparameter, G indicates the actual labels of CT data, and head indicates the segment header, which changes the channel quantity to the segmentation category quantity.
3 Experiment
3.1 Experiment Details
3.1.1 Experiment Environment
Our experiment is performed on MindSpore 1.7.0 and Ascend 910-based hardware. During training, the default batch size is 2, and the SGD optimizer with 0.01 initial learning rate and 0.9 momentum is used to optimize our backpropagation model. In addition, the weights of each loss function for deep supervision are set to 0.53, 0.27, 0.13, 0.07, and 0.
3.1.2 Data Augmentation
For all training, data augmentation polices such as scaling, rotations, brightness, contrast, gamma, and Gaussian noise are used to increase data diversity and robustness to color disturbances.
3.1.3 Patch Sampling
To improve the stability of network training, nnU-Net forces more than one-third of the samples in batches to contain at least one randomly selected foreground class. Due to limited time, the iteration of 50 training batches is defined as an epoch, and a total of 600 epochs are trained. In the end, the 5-fold cross-validation is performed to select an optimal model.
3.2 Experiment Results
According to the results shown in Figure 2, nnU-Net achieves good performance on kidney segmentation (marked in red), but it fails to detect a small number of tumor segmentation parts (marked in green). Furthermore, the last CT image demonstrates an incorrect identification of the tumor beneath the kidney, which could potentially be attributed to inadequate training.
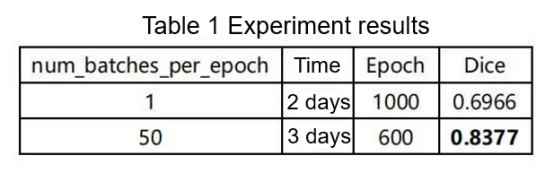
According to the competition organizer, the number of iterations per epoch is 250. Considering limitations in hardware resources and training time, we reduce the number to 50, as shown in Table 1. This is a compromise, as it results in a decrease in accuracy.
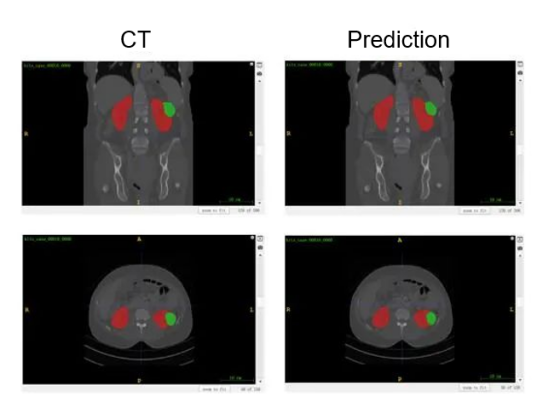
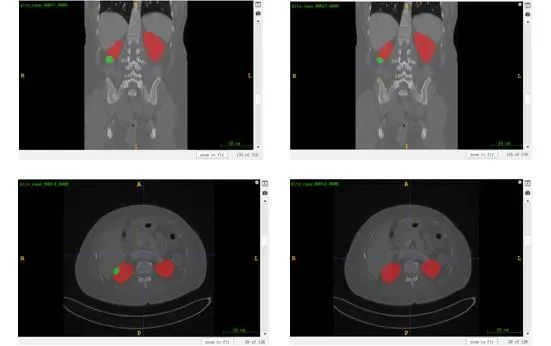
Figure 2 Visualized results of kidney-tumor segmentation
4. Conclusion
When it comes to handling medical data, the challenging aspect lies in comprehending the information and applying appropriate policies for preprocessing, training, inference, and postprocessing. Although classical networks can produce desired outcomes, modifying network structures alone may not result in significant advancements and could potentially cause overfitting issues.
Reference
[1]Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation[C]//International Conference on Medical image computing and computer-assisted intervention. Springer, Cham, 2015: 234-241.
[2]Sen Tao. Researches on deep learning-based kidney-tumor segmentation[D]. Xidian University, 2021.DOI:10.27389/d.cnki.gxadu.2021.001313.
[3]Heller N, Sathianathen N, Kalapara A, et al. The kits19 challenge data: 300 kidney tumor cases with clinical context, ct semantic segmentations, and surgical outcomes[J].arXiv preprint arXiv: 1904.00445, 2019.
[4]Isensee F, Maier-Hein K H. An attempt at beating the 3D U-Net[J]. arXiv preprint arXiv:1908.02182, 2019.
[5]Isensee F, Petersen J, Kohl S A A, et al. nnu-net: Breaking the spell on successful medical image segmentation[J]. arXiv preprint arXiv:1904.08128, 2019, 1(1-8):2.
[6]Isensee F, Jäger P F, Kohl S A A, et al. Automated design of deep learning methods for biomedical image segmentation[J].arXiv preprint arXiv:1904.08128, 2019.